Appyter Workflows for Analyzing Gene Sets with ChEA-KG
The ChEA-KG API has been extended to identify and analyze enriched regulatory subnetworks within two Appyter notebook workflows, which extend Jupyter notebooks to create functional standalone web-based applications. Read more about and try out each Appyter below.
ChEA-KG Appyter
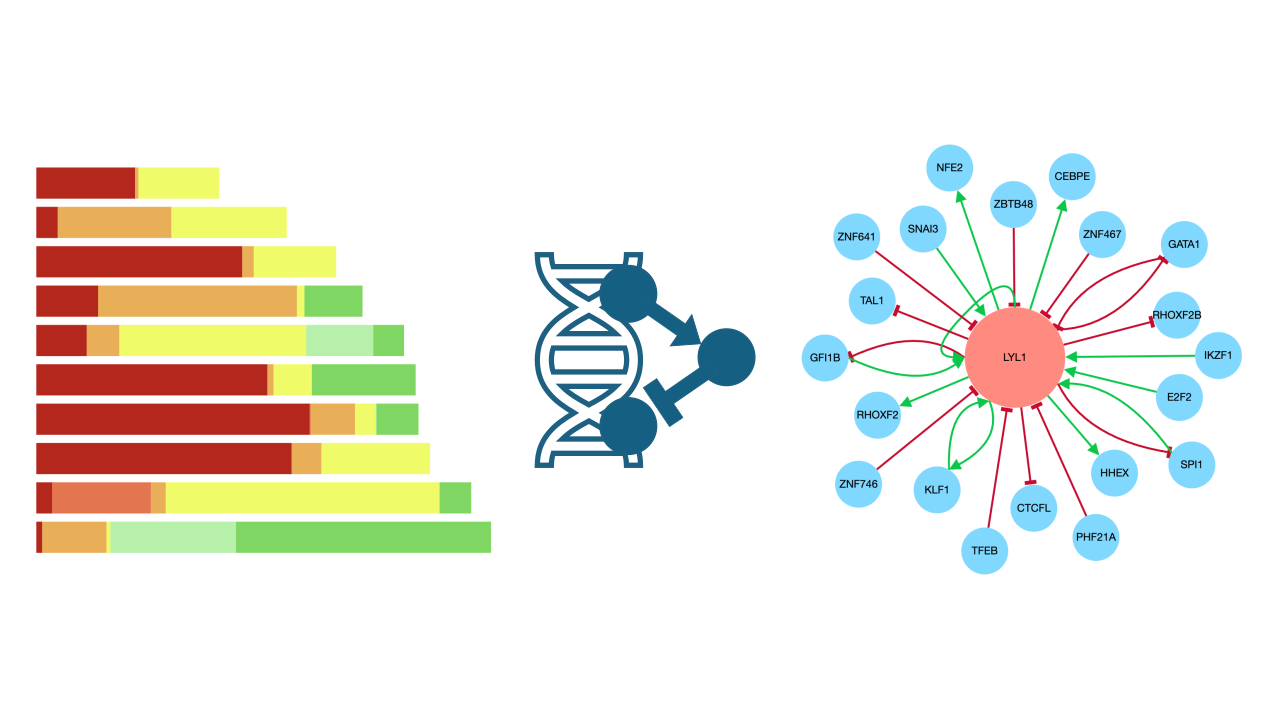
The ChEA-KG Appyter predicts regulatory subnetworks of transcription factors (TFs) for an input gene set. Regulatory networks are generated programmatically using the ChEA-KG API. The workflow then predicts the function of enriched TFs are predicted using the Gene Set Foundation Model.
ChEA-KG-TS
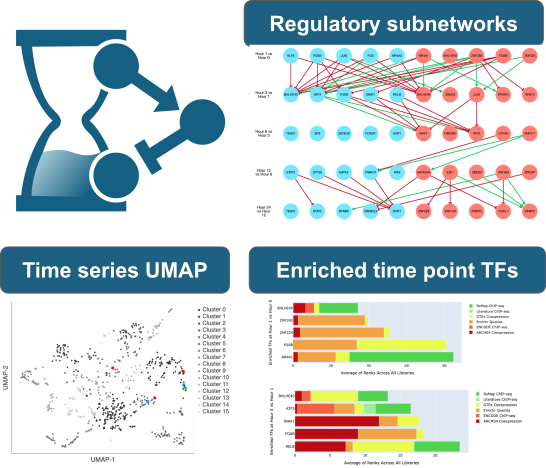
The ChEA-KG Time Series Appyter visualizes the enriched transcription factor landscape from time series RNA-seq data, helping to inform hypothesis about the regulatory mechanisms governing biological processes.

